MAP MAY SURVIVE PASTEURIZATION OF MILK FROM SHEEP
2026-03-22 16:28:41Di Paolo and her Italian colleagues describe experiments designed to assess if MAP can be killed by pasteurization during the production of Pecorino cheese. Their findings were recently reported in the March 14, 2026 issue of the Journal of Food Science. Pecorino romano is a hard, salty Italian cheese made from sheep milk, often used for grating over pasta or other dishes. The name pecorino means “ovine” or “of sheep” in Italian; the name of the cheese, although protected, is a description rather than a brand (Wikipedia).
ABSTRACT
Paratuberculosis is an infectious disease mainly affecting domestic ruminants, caused by Mycobacterium avium subsp. paratuberculosis (MAP). Affected animals shed the pathogen through feces, also during the subclinical stages of the disease. Although there is still no conclusive evidence of MAP involvement in human diseases, particularly Crohn's disease and autoimmune disorders, exposure to MAP through the consumption of contaminated dairy products has become a public health concern. Therefore, in this study, the survival of MAP during the production of Pecorino, an Italian cheese made from raw sheep's milk, was investigated by a microbial challenge test. Three batches of raw milk were artificially contaminated with MAP to obtain a concentration of approximately 104 CFU/mL, higher than natural conditions. The Pecorino cheesemaking process can reduce the initial MAP concentration by about 1 log10 after 60 or 90 days of ripening. The process may act as a hurdle factor against the pathogen, thereby contributing to product safety. Nonetheless, no scientific evidence is currently available regarding MAP contamination levels in raw bulk-tank milk from flocks affected by paratuberculosis, nor quantification of contamination risk relative to flock prevalence. Consequently, as a precautionary measure, milk pasteurization at 72°C for 25 s is recommended to produce Pecorino in flocks with clinical cases of paratuberculosis, regardless of the estimated seroprevalence.
Practical Applications
This study provides relevant insights for artisanal producers of Pecorino and similar cheeses, giving evidence of MAP survival during the cheesemaking process. These findings support product safety assessment and the competent authority for managing potential public health risks.
COMMENT
The research article concludes with this statement as the final sentence of the publication: “Furthermore, given current knowledge, the primary measure to prevent the presence of MAP in sheep milk products is the adoption or implementation of control plans to reduce the prevalence of paratuberculosis within flocks.” This statement is consistent with the main message from authors of this site:
Paratuberculosis control improves animal health and improves producer profitability while simultaneously protecting human health.
Di Paolo's report is one of several studies on the ability of MAP to survive pasteurization and more specifically the various methods of producing cheese. Our website provides much more information on these subjects.
Here are links to pages on our site for those interested in learning more:
... about heat tolerance of MAP: https://johnes.org/resistance/
... about finding MAP in food and water: https://johnes.org/is-map-in-food-and-water/
... and, for lectures on MAP and human health: https://johnes.org/presentations-and-mini-lectures/
IN UTERO TRANSMISSION OF MAP IN SAANEN GOATS.
2026-03-01 22:48:03
G.V. Sandoval and colleagues in Argentina reported on the rate of intrauterine MAP transmission in pregnant Saanen goats.
Their findings were reported in a recent issue of the journal Veterinary Microbiology.
ABSTRACT
Paratuberculosis (PTBC) is a chronic granulomatous enteritis of ruminants caused by Mycobacterium avium subsp. paratuberculosis (Map). Although PTBC intrauterine transmission has been described in cattle, sheep, and deer, data in dairy goats are limited. The present study investigated intrauterine Map infection in 32 pregnant Saanen goats from a herd with high PTBC prevalence in Jujuy, Argentina. Maternal and fetal tissues were analyzed using a range of methods, including bacteriological culture, IS900 PCR, histopathology, and serology. All goats were positive to the intradermal test, and 75% showed histopathological lesions consistent with PTBC. Map was isolated from 10 of 45 fetuses. Granulomatous lesions were observed in two cases, but no acid-fast bacilli were detected. Furthermore, fetuses from culture-positive dams were 7.8 times more likely to be infected than those from culture-negative ones. This suggests that the maternal bacterial load is a key predictor of fetal infection. These findings represent the first evidence of intrauterine transmission of Map in Saanen goats, even in subclinical cases, indicating that vertical transmission may be more common in this species than previously assumed. This has significant implications for PTBC control programs in small ruminants.
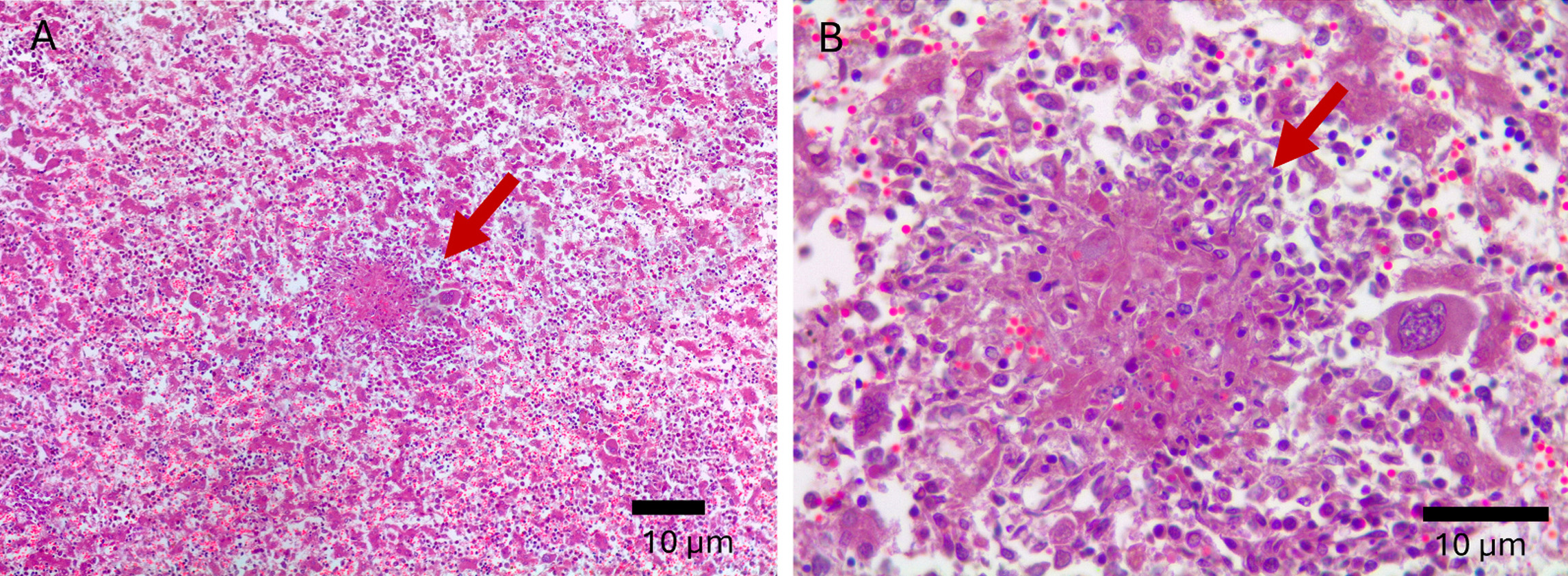
Liver granulomas from MAP-infected female goats indicating they had a disseminated infection. H&E stain. A. 10x magnification B. 40x magnification. Sandoval et al. Veterinary Microbiology 315 (2026) 110954.
COMMENTS
This intriguing study that verifies that MAP infections can pass from mother (dam) to off-spring before birth, i.e., in utero. The best predictors of whether in utero transmission would occur were culture of maternal tissues collected at necropsy and PCR on these same tissues. Interestingly, ELISA on the dams did not predict the likelihood fetal infection, i.e. there was no statistical association between the ELISA-positive status of the pregnant goat and the MAP infection status of its fetus.
Why this study might not represent all goat herd situations:
- This study was done on a single heavily MAP-infected herd of Saanen goats.
- The herd had a very high MAP infection rate, estimated to be 23%. This means that the female goats in the study were likely exposed to very high numbers of MAP causing more rapid infection progression. These females themselves may have become MAP-infected in utero.
Take home messages for herd owners working to control or eradicate Johne’s disease:
- PCR is preferable to ELISA for herd testing, especially for breeders.
- It is best to not keep as herd replacements any kids born to PCR-positive dams.
- Owners cannot simply trust their eyes to diagnose MAP-infected goats. All dams in this study were in good body condition (body condition score over 3), but 14 of 32 had advanced MAP infections based on histopathology. Weight loss and diarrhea do not happen until very late in the course of a MAP infection in goats. Thus, goat owners must rely on PCR on fecal samples to detect MAP-infected goats, although this study did not report fecal PCR test results on live goats but instead relied on PCR on tissue samples collected at necropsy.
JD CONTROL IN BEEF HERDS IS COST-EFFECTIVE
2025-12-21 16:41:12Veterinarians and economists at the University of Saskatoon in Canada teamed up to evaluate the cost-effectiveness of Johne’s disease (JD) control strategies in beef cattle herds. They concluded that “testing every year with individual PCR provided the best mix of disease reduction and financial return.”
This scientific article was published December 17, 2025 in the journal Veterinary Sciences by L. Rosengren et al. It is very technical and best suited for experts. However, the main message is clear, consistent with most other such studies, and in full agreement with the recommendations presented for beef cattle herds on this website.

As stated in the article:
“The primary objective of this study was to evaluate test-and-cull strategies for Johne’s disease in Western Canadian beef herds using an existing stochastic simulation model. Specifically, strategies were assessed against three criteria: (i) feasibility within typical animal handling patterns, (ii) effectiveness in maintaining or reducing JD prevalence at 10 years, and (iii) profitability compared with no testing, as measured by the 10-year NPV. A secondary objective was to quantify the relative economic cost of Johne’s disease to the beef industry and to assess the value of maintaining JD-free herds under different replacement and market scenarios.”
SIMPLE SUMMARY
Johne’s disease (JD) is an untreatable intestinal infection in cattle that reduces herd health and profitability. To understand how well different control programs work in beef herds, we combined a computer simulation to model JD spread in a typical 300-cow Western Canadian operation with an economic model to estimate financial outcomes over 10 years. We evaluated seven test-and-cull strategies that used different diagnostic tests and testing schedules, including risk-based sampling. All strategies lowered the frequency of JD compared with doing nothing, and most improved overall profitability. Testing every year with individual PCR provided the best mix of disease reduction and financial return. Testing every six months with PCR reduced disease the most but cost more. Herds that did not use any control saw JD levels rise and experienced long-term economic losses. The results were generally stable across changes in production costs, market prices, and replacement-heifer management. However, profits dropped when purchased animals had a high JD prevalence. Overall, this study shows that JD can be economically controlled in beef herds using long-term test-and-cull programs tailored to producer goals.
COMMENTS
The findings of this study are generally applicable to other types of animals affected by JD, especially for animals used for breeding purposes.
For general information about JD in beef cattle visit this page of our website.
To watch a one-hour lecture on JD in beef cattle click here.
NATIONAL CONTROL PROGRAMS CAN WORK
2024-12-05 16:06:40Great Britain has significantly decreased the rate of MAP infections among the country’s dairy herds. This ongoing effort is succeeding because of the involvement of dairy producers, veterinarians, dairy manufacturers, and governmental animal health authorities. The result is improved animal health, improved farm economics and efficiency, and improved food safety for consumers. A detailed analysis of data spanning 10 years has been recently reported in the journal Preventive Veterinary Medicine.
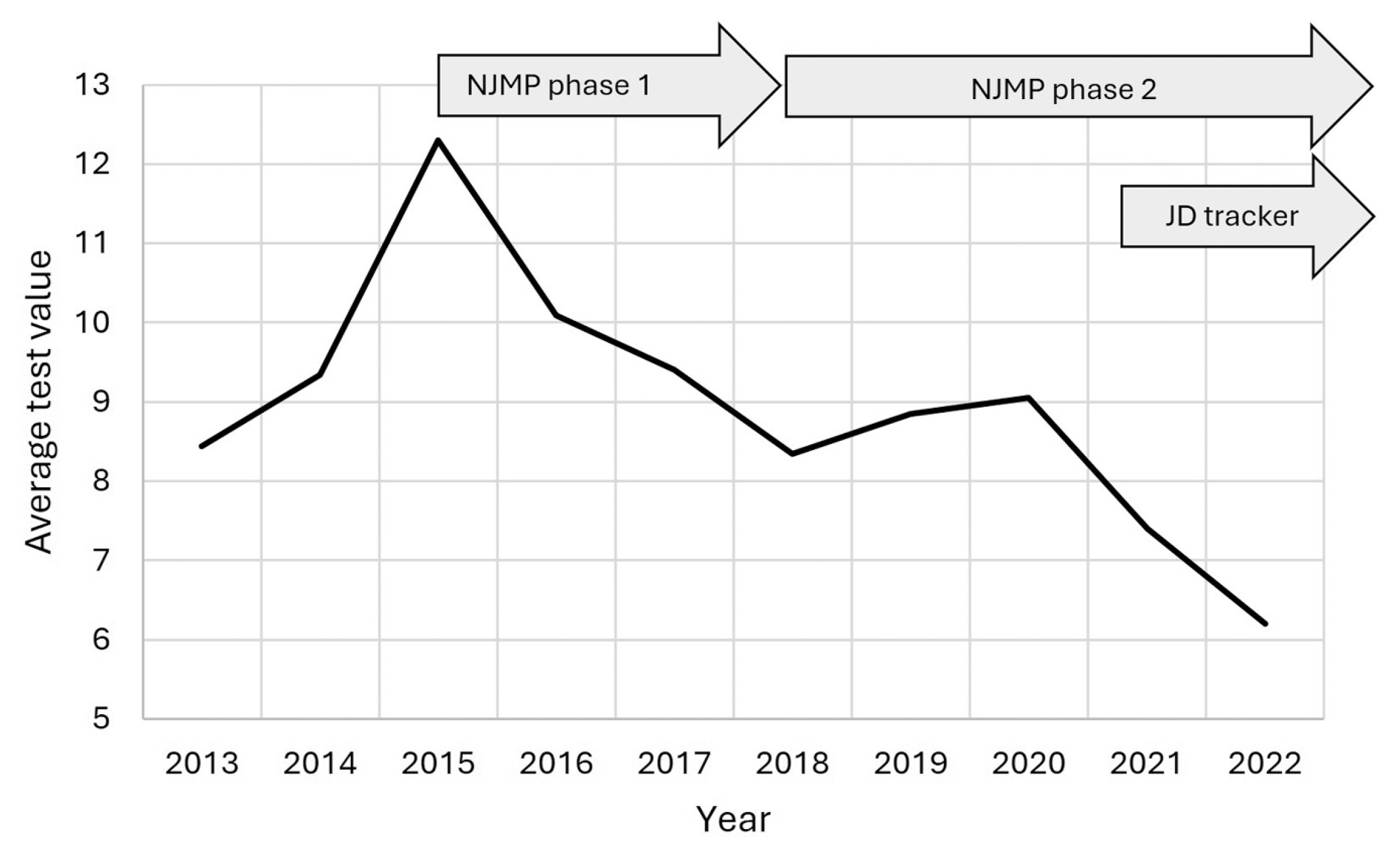
Figure: The pooled ATV (average test, meaning milk ELISA, value) for JD (Johne’s disease) tests carried out during ‘whole herd’ tests in the 154 studied herds, by year. NJMP Phase 1 and 2 refers to the National Johne’s Management Plan Phases 1 and 2.
The article is written for technical specialists and has an unusually high number of acronyms to keep straight. However, by all the many measures used to track MAP infections, dairy herds in Great Britain have fewer MAP-infected dairy cattle today than they did a decade ago.
ABSTRACT
Effective management of cattle infected with Johne’s Disease (JD) is crucial to minimizing transmission and within-herd prevalence. Within Great Britain (GB), the voluntary National Johne’s Management Plan (NJMP) requires farmers and a certified vet to conduct a risk assessment to determine the herd risk, examine the herd JD status and formulate a management plan. Individual milk ELISA tests for JD antibodies are widely used to monitor infection. The JD Tracker application, available within the dairy data management software InterHerd+ and other web-based environments, is being used by farmers and veterinarians to facilitate the practical use of milk ELISA data to aid JD-related management decisions. The JD Tracker application uses a herd’s milk ELISA data to calculate a collection of ‘JD parameters’ that are indicative of the current JD status of the herd alongside contemporary and retrospective drivers linked to transmission and maintenance of infection. Herein, we use milk ELISA data from 154 regularly testing herds to review the temporal trends in JD parameters from 2013 to 2022. Since 2015, JD Tracker parameters have improved in these herds, most notably average test value (ATV) and within-herd prevalence (%Pos30). Trends in driver parameters suggest that farmers are progressively less likely to serve repeat test-positive (J5) cows and are more readily removing them. The data also reveal that the burden of JD is disproportionately greater in herds with higher ATV. In 2022, the 25 % of herds with the highest ATVs accounted for 42 % of positive tests and 42 % of repeat ELISA positive (J5) cows. Retrospectively, it is not possible to identify with certainty factors that directly contributed to the trends in JD parameters, but it is notable that the introduction of the NJMP was coincided with the improving JD situation. In 2019, participation in the NJMP or an equivalent scheme became mandatory for dairy farms to be compliant with the food and farms standards assurance scheme Red Tractor, with the result that JD management plans are now completed by 95 % of UK dairy farms. As far as we know, the UK is unique in its development of a tool (the JD Tracker) which adds utility to milk ELISA data using specifically designed JD parameters. Anticipated further work includes the development of a national database of JD testing herds and application of the JD Tracker at national scale to enable more comprehensive industry-level monitoring of JD within GB dairy farms.
COMMENTS
This article demonstrates that Johne’s disease control is achievable at the national level when stakeholders at every level of the supply chain, including consumers (taxpayers), are engaged. Programs that depend too heavily on governmental funding often are unsustainable because of the long-term necessity of continued financial support in the face of declining budgets for animal health.
CROHN’S CURE?
2024-08-03 15:59:30Dr. David Graham from Baylor College of Medicine together with 16 international colleagues published the results of a large trial to treat Crohn’s disease with anti-MAP antibiotics. This study met the highest standards for a clinical trial, namely it was a multi-center randomized, double-blind, placebo-controlled study. This clinical trial was conducted at 92 sites in the United States, Canada, Bulgaria, the Czech Republic, Australia, New Zealand, Israel, Poland, Serbia, and Slovakia. The article was published in the journal Antibiotics and is Open Access (free for everyone).
ABSTRACT
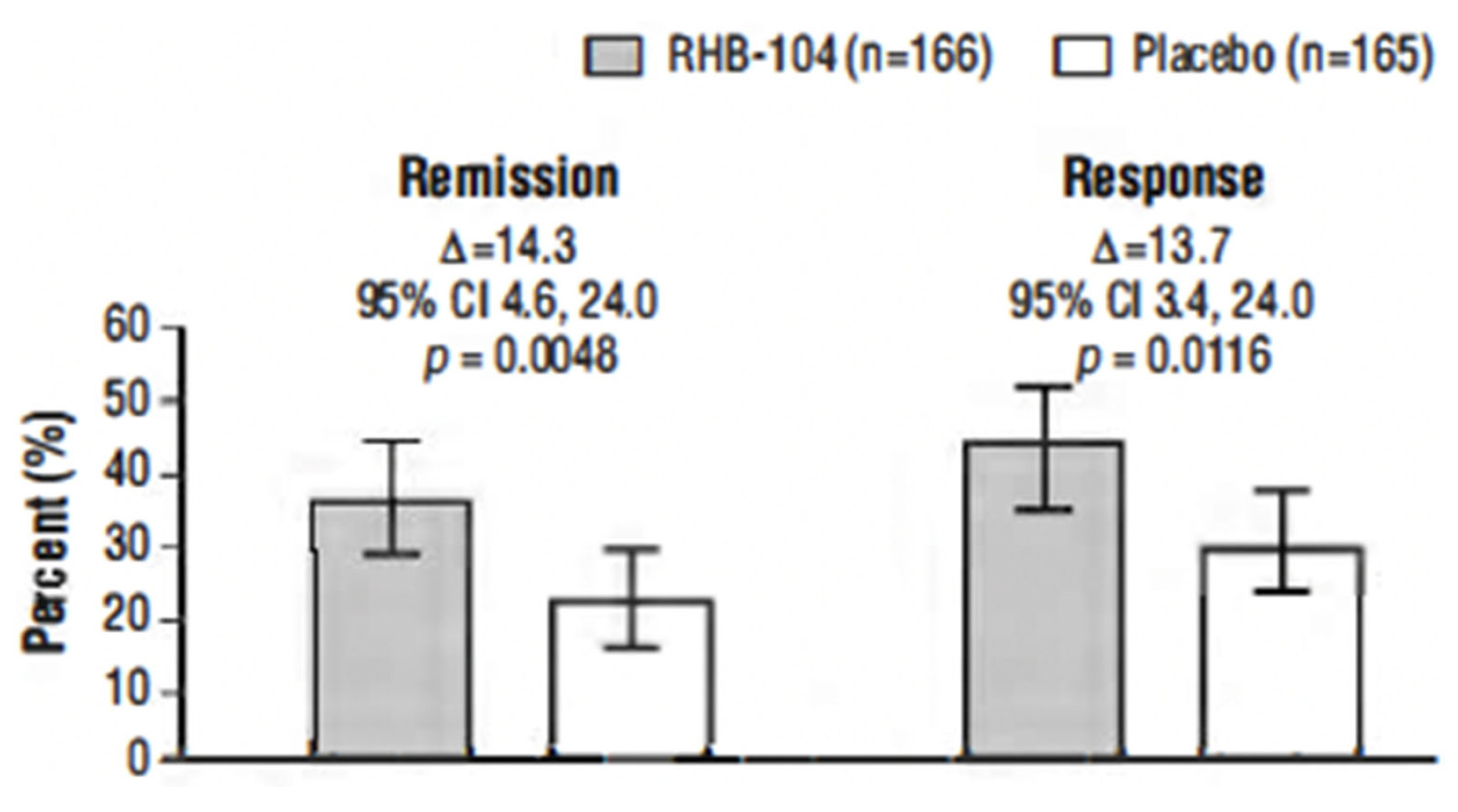
This study, conducted between 4 October 2013, and 30 November 2018, tested the hypothesis that triple antimicrobial therapy, targeting Mycobacterium avium subspecies paratuberculosis (MAP), long considered a putative cause, would favorably affect Crohn’s disease. A double-blind multicenter study of adults with active Crohn’s disease, (i.e., Crohn’s Disease Activity Index [CDAI] 220–450 plus C-reactive protein ≥ 1.0 mg/dL, fecal calprotectin (FCP) >162.9 μg/g stool, or recent endoscopic or radiographic confirmation of active disease) receiving concomitant standard-of-care Crohn’s disease treatment (Clinicaltrials.gov: NCT01951326) were stratified by anti-tumor necrosis factor use and randomized (1:1) to anti-MAP RHB-104 (clarithromycin 95 mg, rifabutin 45 mg, and clofazimine 10 mg per capsule) (n = 166), resulting in clarithromycin 950 mg/day, rifabutin 450 mg/day, and clofazimine 100 mg/day, or placebo (n = 165) for up to 52 weeks. A greater proportion of RHB-104 versus placebo-treated patients met the primary endpoint—remission (i.e., CDAI < 150)—at week 26 (36.7% [61/166] vs. 22.4% [37/165], respectively; 95% CI for difference: 4.6, 24.0, p = 0.0048; chi-square test). Clinical response (reduction of CDAI by ≥100 points from baseline) at week 26 (first secondary endpoint) was also higher among the patients treated with RHB-104 (73/166 [44.0%]) compared with placebo (50/165 [30.3%]; 95% CI for difference: 3.4, 24.0, p = 0.0116), and it remained higher at week 52 among the patients treated with RHB-104 (59/166 [35.5%] vs. (35/165 [21.2%] for placebo; 95% CI for difference: 4.7, 23.9, p = 0.0042). A statistically significantly greater decline in FCP (another prospective efficacy endpoint) was also observed in RHB-104-treated patients, compared with placebo, at weeks 12, 26, and 52. The rates of serious adverse events were similar between groups (RHB-104: 18.7%; placebo: 18.8%). No patient died during the study. Antimicrobial therapy directed against MAP resulted in significantly greater improvement in clinical and laboratory (FCP) measures of active Crohn’s disease.
COMMENTS
There have been multiple scientific reports on curing individual cases of Crohn’s disease using anti-MAP antibiotics. The article in the journal Antibiotics is the largest clinical trial testing a treatment for Crohn’s disease directed at a cause of Crohn’s disease, namely MAP, rather than at the symptoms of the disease, e.g. immunosuppressive drugs. It significantly adds to the body of evidence that MAP is a zoonotic bacterial pathogen causing chronic intestinal inflammation in a diverse array of animal species.
For a true story of a veterinarian permanently cured of his Crohn’s disease, and links to similar case reports, visit this page of the website: https://johnes.org/other-animals/non-ruminants/
For more on the zoonotic potential of MAP, and links to scientific articles on the subject, visit this page of the website: https://johnes.org/zoonotic-potential/.
For more on MAP in food and water visit this page: https://johnes.org/is-map-in-food-and-water/.
For a comprehensive 90-minute lecture on scientific evidence that MAP is a cause of Crohn’s disease see this page: https://johnes.org/presentations-and-mini-lectures/
MAP-CROHN'S CONUNDRUM
2024-07-12 16:49:39Dr. Robert J. Greenstein has published an article titled: Human genetic defects and misinterpreted pharmacological data indicate that Crohn disease is consequent to a mycobacterial infection in the medical journal, Medical Research Archives, published by The European Society of Medicine. The Open Access article (8 pages with 81 references) appears in the soon to be published volume 12, issue 7 of the journal. It is somewhat technical, but the underlying premise is readily understood by non-experts and should be of particular interest to those concerned about the zoonotic potential of MAP.
ABSTRACT
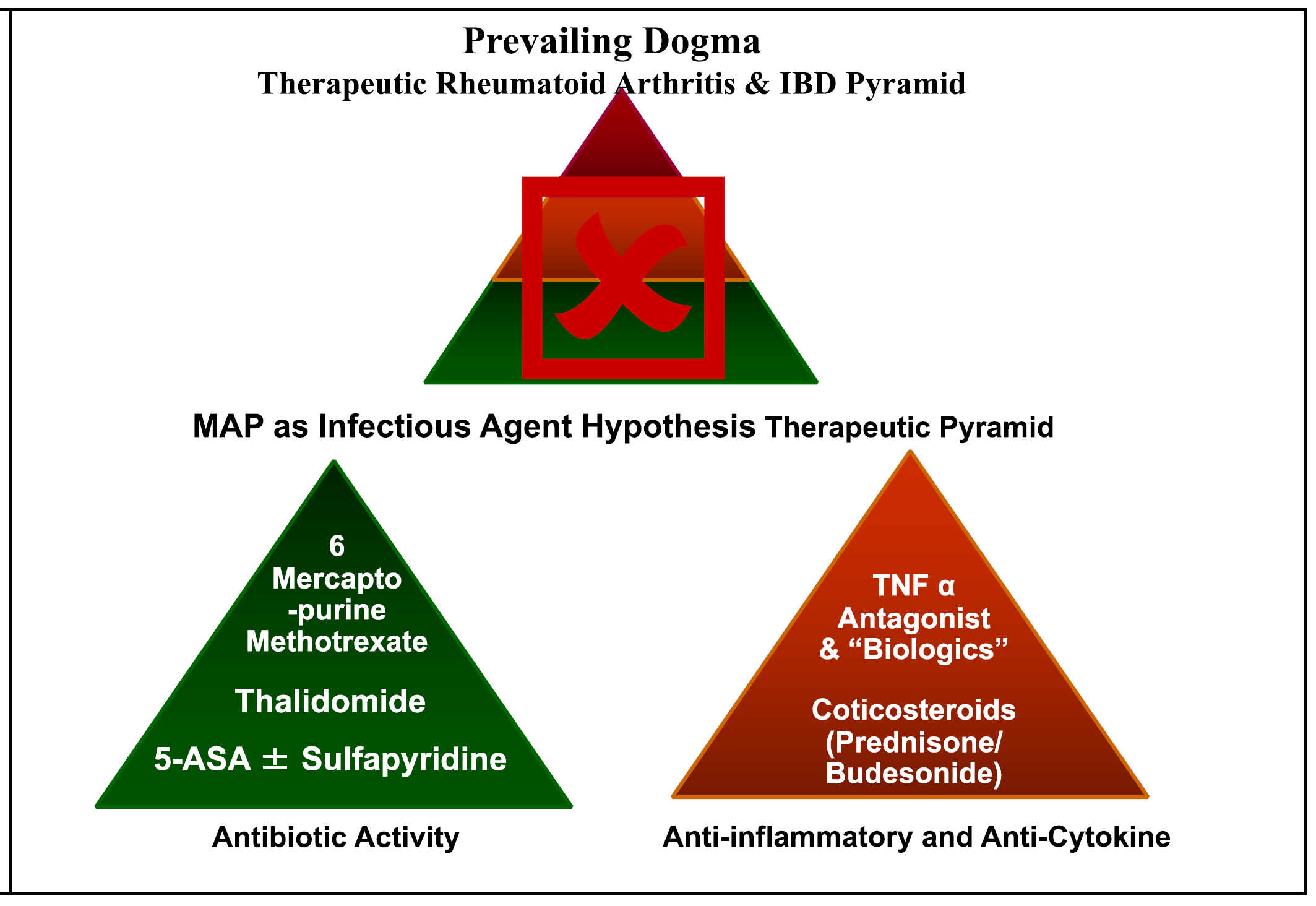
For more than a century, there has been controversy whether Crohn's disease is a cryptic zoonotic mycobacterial infection. In this manuscript we address two, usually ignored, critical elements that suggest that this concern is probably correct. First: human genetic defects identified in Crohn's disease are associated with increased susceptibility to mycobacterial infections. Second: multiple pharmaceutical agents that are used in Crohn's disease are misnamed. Many "anti-inflammatories" and "immune modulator" medications have dose dependent inhibition of mycobacteria in culture. Accordingly, their primary mechanism of action are as antimycobacterial antibiotics. Failure to incorporate these data renders all previous antibiotic studies in Crohn's disease irrevocably flawed. Finally, incorporating these genetic and antibiotic data, we suggest the necessary clinical studies to address the hypothesis that Crohn's disease is a zoonotic infection.
CONTROL PROGRAMS FOR DAIRY HERDS
2024-05-01 18:33:32An international team of researchers, led by Maarten F. Weber from The Netherlands, has published a paper summarizing the state of national paratuberculosis control programs for dairy cattle herds across the globe with a focus on the most important drivers of program success. The publication is in the April issue of the Open Access journal Animals.
This well-written article gives historical background and summarizes the many lessons learned by programs in multiple countries. While primarily focused on dairy cattle, there is mention of goats and sheep, and the principles of paratuberculosis control apply broadly. The content is readily understood by non-scientists and the 96 cited references lead interested readers to the primary literature. This article describes how concerted efforts at the national level can lead to successful eradication of paratuberculosis, as was done in Norway and Sweden. It also provides an excellent summary of the most effective control measures at both the national and farm levels.

SUMMARY (British spellings)
Paratuberculosis control programmes in countries with a relevant dairy industry differ largely in participation and progress. Despite over a century of experience with paratuberculosis control efforts, major knowledge gaps still exist, including the efficacy of control programmes and drivers and barriers influencing the uptake of control programmes amongst farmers. Biennially, the International Dairy Federation (IDF) brings together experts on paratuberculosis control to share the most recent knowledge and experiences regarding practical aspects of paratuberculosis control.
Taken together, studies on control programmes presented at the 7th and 8th IDF ParaTB Fora and the 15th International Colloquium on Paratuberculosis (ICP) indicated a key finding that a reduction of the prevalence of Map infection had been achieved by various programmes. Important prerequisites for successful control were long-term stable funding, stakeholder commitment and incentives for farmers to participate. Focal topics to improve the control of Map were identified, including improved communication about the epidemiology of infection and its control, increased attention to intrauterine, calf-to-calf and adult-to-adult transmission, sound but easy-to-use surveillance schemes, measures to reduce between-herd transmission and breeding for resistance to Map infection. Research in parallel with these programmes was found to keep interest in Map control high among stakeholders and farmers and to enable programme improvement.
COMMENTS
The key findings of this publication bear repeating for emphasis.
The success of national control programs requires:
- Long-term, stable funding.
- Education of producers and veterinarians.
- Veterinary risk assessments, control program design and management on farms.
- Commitment from stakeholders, especially milk processors. As stated in the article: “North American milk processors did not buy into the programmes, leaving participating farmers and veterinarians without incentives to participate.”
- Ongoing collaborative and practical research.
- Financial incentives for dairy farmers. Farmers cannot shoulder the full cost of paratuberculosis control, and they shouldn’t, because of the probable human health/societal impacts of this infectious, zoonotic disease.
MAP: A PROBABLE CAUSE OF CROHN’S DISEASE
2024-04-22 16:22:32
For the past 10 years I have been invited to give a lecture in a public health and epidemiology course, PHS-801, at the University of Wisconsin-Madison, School of Medicine, and Public Health as part of a series of lectures on disease causation, i.e., judging scientific evidence as to whether a pathogen is the cause of a disease. Every year the lecture was refined and updated with new information. On April 9, 2024, I again gave the lecture and this year had it recorded live. The 80-minute lecture is now posted on the Presentations page of my website: https://johnes.org/presentations-and-mini-lectures/ (scroll down to the last entry).
The lecture cites numerous peer reviewed scientific publications and in places is very technical. Most of those publications can be found on the page titled Zoonotic Potential: https://johnes.org/zoonotic-potential/. However, many parts of the lecture are quite understandable to non-scientists as the goal is to provide a “big picture” perspective on this global problem regarding MAP as a probable human pathogen.
SUCCESSFUL ERADICATION FROM A DAIRY HERD
2024-04-08 15:56:45K. Donat and colleagues reported on the successful control and likely eradication of MAP from a 450-cow commercial dairy herd in Germany. This case study, lasting a decade (2012-2022), was published in the Open Access journal Animals March 21, 2024.
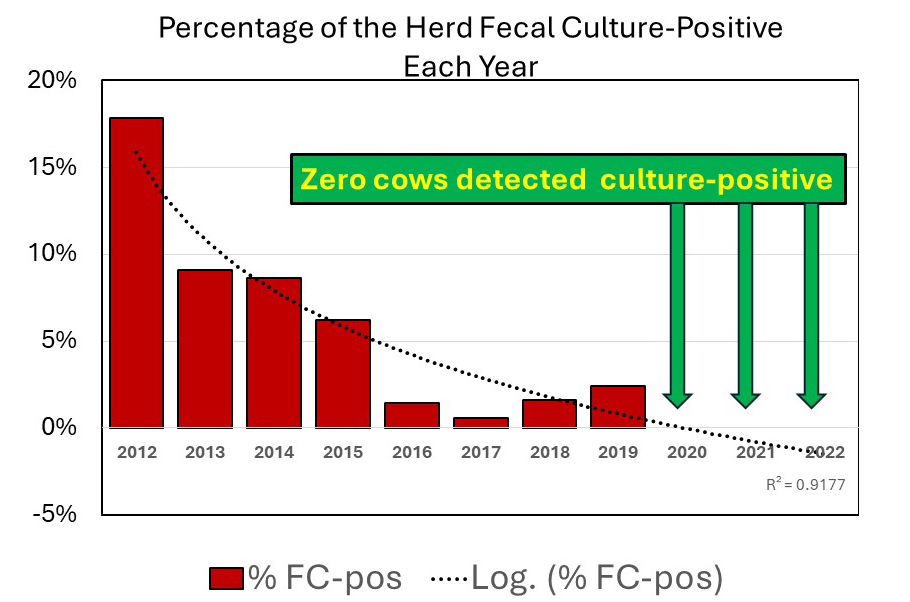
The above graph was generated from the study data provided in Table 1 of the publication (red bars) with the addition of a logarithmic regression line (dotted black line). The herd had a high infection rate in 2012 with 17.83% of cows being fecal culture-positive and from 2020 to 2022 the herd had zero culture-positive cows.
ABSTRACT
This longitudinal case study provides an in-detail report of the process towards the elimination of Mycobacterium avium subsp. paratuberculosis (MAP) from a closed 450-head commercial dairy herd. In parallel, two diagnostic approaches were applied to all cows in annual intervals during 2012–2022: detection of MAP in individual faecal samples by bacteriological cultivation on solid medium and detection of MAP-specific antibodies by ELISA. For each annual sampling, the kappa coefficients for test agreement and the survival rates of MAP-positive and MAP-negative cows were calculated. Applying a multivariable linear regression model revealed a significantly lower fat-corrected 305-day milk yield for MAP-positive cows. The true prevalence of MAP shedders reduced from 24.2% in 2012 to 0.4% in 2019 and during 2020–2022, no MAP shedder was identified. Test agreement was generally low and bacteriological cultivation showed positive results earlier than the ELISA. In the first years of control, the survival of MAP shedders was longer than in the final stage. In conclusion, the elimination of MAP from a dairy herd might be feasible within a decade. Changes in the test agreement must be considered. Timely removal of MAP shedders, hygienic calf rearing, and colostrum supply are key for successful control.
COMMENTS
This publication is worth reading in detail, especially for dairy producers and veterinarians focused on dairy cattle. The basic principles also apply to other animal species. This case study contains many useful observations. Here is just one example: “….the farmer observed better fitness in multiparous cows and a more pronounced drop in milk of MAP shedders of second and higher parity. This was worth spending EUR 57,963 (US $62,887) on paratuberculosis control during 11 years, which is approximately EUR 11.70 (US $12.69) per cow per year.”
There are several important lessons from this case study and here is my list of important lessons, based both on this publication and others like it:
- Paratuberculosis control takes time and therefore patience.
- Farmer engagement and continuous support is vital for success.
- Veterinary supervision and monitoring are a critical, and often overlooked, part of the program.
- Herd management changes are essential for effective paratuberculosis control.
- Regular herd testing to identify the most infectious cows for segregation at calving time and then culling is necessary.
- Governmental programs with funding to offset the costs of paratuberculosis control help make control programs financially feasible for producers.
- Paratuberculosis eradication can be achieved.
SICILY: SHEEP & GOAT SURVEY
2024-02-25 17:36:43 Vincenzo Di Marco Lo Presti and 11 colleagues from Italy, Ireland, and the UK reported results of their large-scale survey of sheep flocks and goat herds (48,643 animals in 439 flocks or herds) in Sicily for evidence of MAP infections using a commercial ELISA kit. Their findings were reported in the February issue of Frontiers in Veterinary Science (Open Access).
Vincenzo Di Marco Lo Presti and 11 colleagues from Italy, Ireland, and the UK reported results of their large-scale survey of sheep flocks and goat herds (48,643 animals in 439 flocks or herds) in Sicily for evidence of MAP infections using a commercial ELISA kit. Their findings were reported in the February issue of Frontiers in Veterinary Science (Open Access).
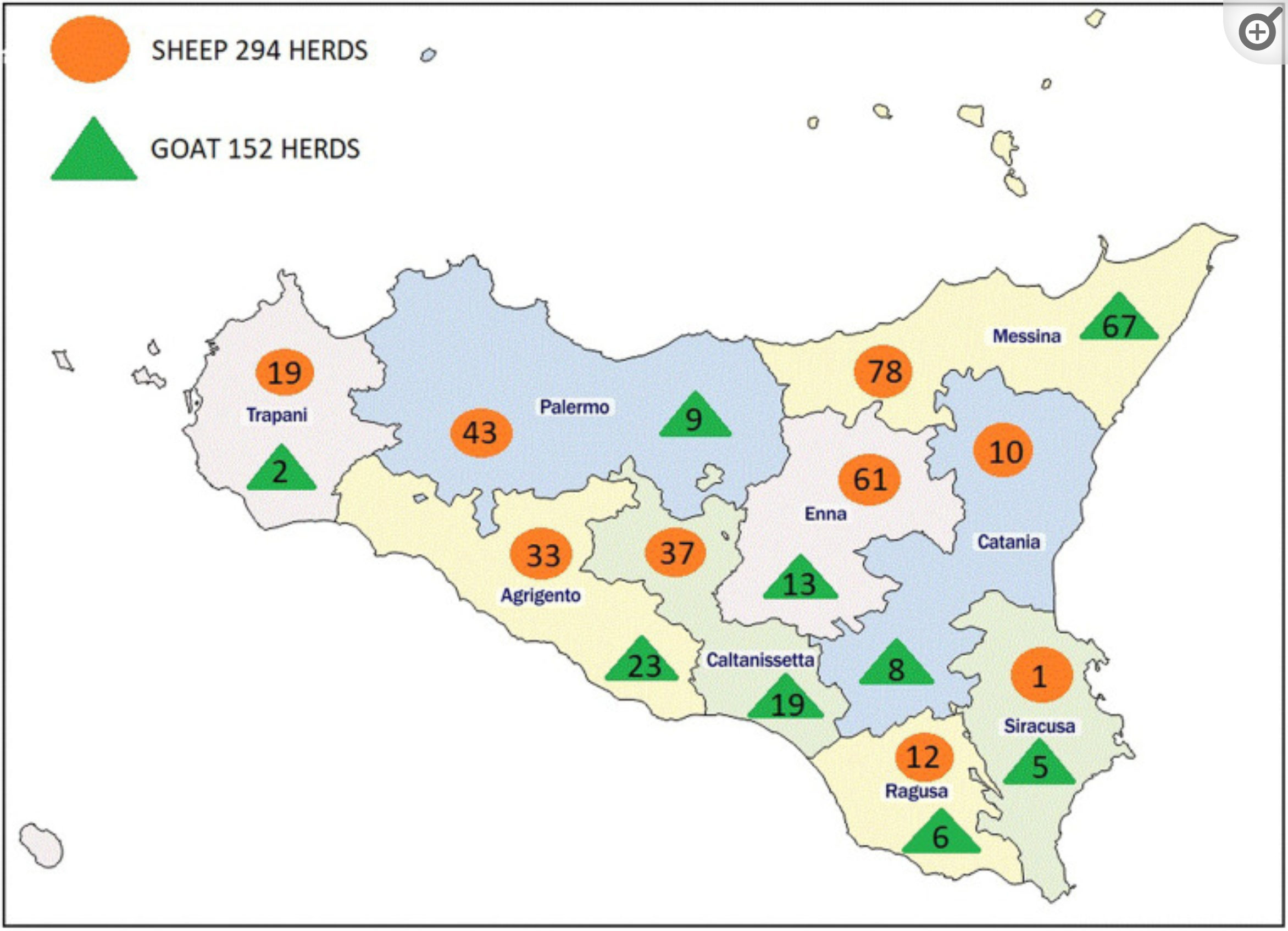
Location and number of herds and flocks tested in Sicily (from publication).
ABSTRACT
Introduction: Paratuberculosis (PTB) is a worldwide chronic, contagious enteric disease caused by Mycobacterium avium subsp. paratuberculosis (MAP) mainly affecting ruminant species. PTB is a WOAH-listed disease with direct and indirect economic losses in the livestock sector, negative impact on animal welfare and significant public health concerns. In spite of this, MAP prevalence in small ruminants is still unknown and the prevalence appears to be underestimated in many countries. The aim of this study is providing a first large-scale serological survey on MAP infection in small ruminants in Sicily, a region of Southern Italy with the 11.3 and 8.9% Italian national heritage of sheep and goats, respectively.
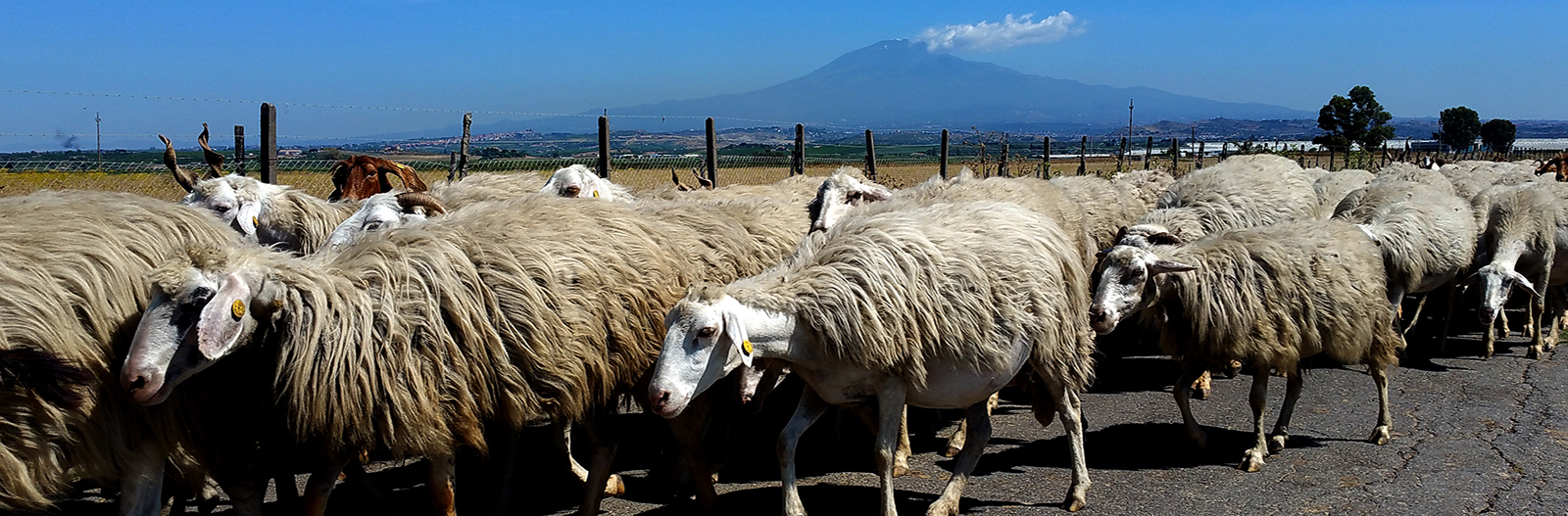

Sicilian landscape with flock of sheep in the foreground and the active volcano, Etna, in the background.
Methods: For this purpose, we analyzed a total of 48,643 animals reared in 439 flocks throughout Sicily. MAP seroprevalence was estimated both at herd-level and animal-level within breeds reared in all the nine sampled provinces.
Results: Our results revealed a high overall apparent prevalence at herd-level of 71.8% in sheep and 60.8% in goat farms with an animal-level prevalence of 4.5 and 5.1% in sheep and goats, respectively. Significant statistical differences were found between the provinces and within the breeds both in sheep and goats.
Discussion: Our study provides the first large-scale serological survey on PTB infection in small ruminants in Sicily and showed a high prevalence of disease depending on the species, breed and province. This study represents the first step to better understand the MAP epidemiology in a typical Mediterranean breeding context, suggesting the need of in-depth study on the herds risk factors, including the eventual presence of candidate genes for resistance/susceptibility to PTB in native breeds.
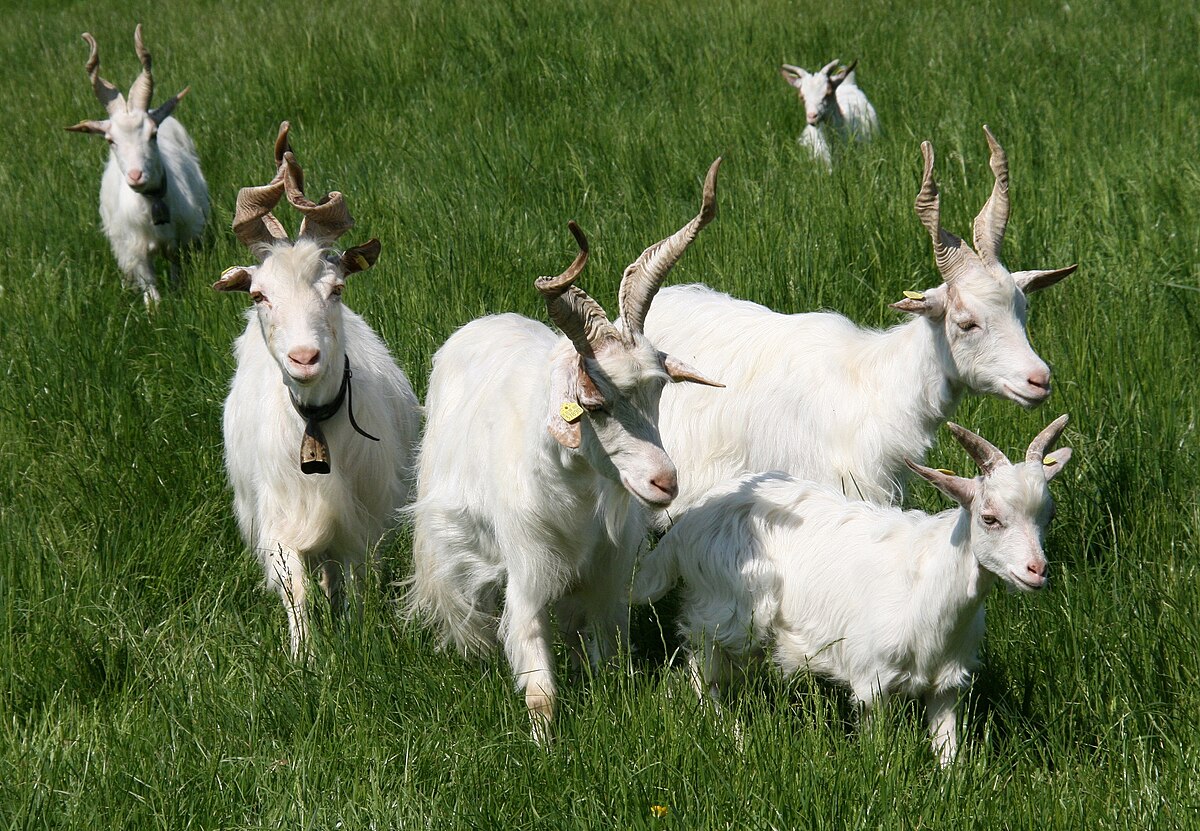

Girgentana goats.
COMMENTS
This large, well-designed and analyzed serological survey confirms what many similar, but smaller, studies have shown: MAP infections are common in sheep and goat populations worldwide. MAP infections threaten valuable breeding stock, such as the Girgentana goats and Barbaresca Siciliana sheep, and local wildlife. Meat and milk products made from MAP-infected animals can be contaminated with live MAP which are resistant to food manufacturing practices and pose a probable health risk for humans. Much more effort and investment are required to control this prevalent infection globally.

This website has much more information specifically about Johne's disease in goats and sheep - - just a click away.
1 2 3 … 18 Next »
